Main Area of Focus
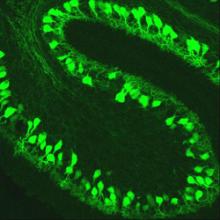
Gene Expression Nervous System Atlas (GENSAT) is an NIH-funded, publicly available gene expression atlas of the developing and adult central nervous system in the mouse. The project also provides transgenic BAC-EGFP reporter and BAC-Cre recombinase driver mouse lines, which are available as research resources to the scientific community.
Background
The NIH-sponsored GENSAT (Gene Expression Nervous System Atlas) project has created close to 1,500 transgenic mouse lines expressing green fluorescent protein (GFP) reporters or DNA recombinases in specific neural and glial cell populations. In each mouse line, expression of the reporter or recombinase is controlled by promoter elements derived from a bacterial artificial chromosome (BAC) containing a specific gene of interest, in order to mimic the gene’s expression pattern. Many of these lines have proven to be extremely valuable in experiments requiring identification of specific cell populations and details of cellular morphology. Furthermore, each of these lines is a unique reagent that provides important data regarding the potential utility of its specific BAC targeting vector. In collaboration with the NIMH Intramural Program, GENSAT has also generated BAC-Cre recombinase driver lines to serve as tools for cell-specific genetic manipulations in the CNS. Over 200 fully characterized BAC-Cre recombinase lines have been created, targeting selected neuronal or glial populations in the brain and spinal cord (also see PMID: 17855595).
The BAC-GFP expression data and mouse brain images are available to the public in on-line, searchable databases (see weblinks below). Since the transgenic BAC mouse lines are powerful tools for neuroscience research, the NIH Blueprint supports the distribution of GENSAT mouse strains via the Mutant Mouse Regional Resource Center (MMRRC). Over 1100 BAC-GFP reporter mouse lines and over 275 BAC-Cre recombinase driver lines have been placed in the MMRRC repositories and are available for a small processing fee.
Contacts
Dr. Laura Mamounas, Ph.D.
Program Director
NINDS
301-496-5745
Related Resources
- BAC Transgenic Mouse GENSAT Database
- Mutant Mouse Regional Resource Centers (using ‘advanced search form,’ select major collection ‘NINDS/GENSAT’ and mutation type ‘Transgenic (cre)’ or ‘Transgenic (BAC)’)
