Main Area of Focus
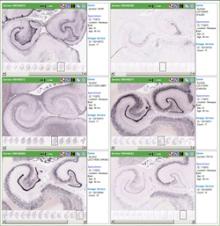
The Allen Institute for Brain Science (Seattle, Washington) was contracted to conduct a high throughput analysis of gene expression in the developing Rhesus macaque brain. Under a subcontract, the University of California Davis provided Rhesus macaque brain tissue for gene expression analysis.
A publicly accessible gene expression map of the macaque brain is available. This project serves as an important link between efforts to map gene activity in the mouse brain and similar efforts to map the human brain.
Background
The NIH Blueprint NHP atlas is an online database of gene expression in the rhesus macaque brain from 6 prenatal stages and four postnatal stages including birth to four years old. The atlas is publicly accessible and allows users to search for gene expression data by gene, brain region, and developmental stage.
The atlas serves as a valuable reference tool for developmental neuroscience research. Among other things, it allows researchers to examine how different genes and gene networks affect primate brain development, and to identify potential genetic targets for therapy in developmental disorders. In combination with similar efforts to map gene expression in the mouse and human brain, it enables researchers to explore what makes our brains unique.
The atlas focuses on five brain regions implicated in a variety of human neurodevelopmental disorders, including prefrontal cortex, primary visual cortex, hippocampus, amygdala and the ventral striatum. Data were collected at ten timepoints: 6 prenatal (E40, E50, E70, E80, E90 and E120) and 4 postnatal (birth, infant -3 months, juvenile-1 year and young adult-4 years) stages. The data include:
- Cellular resolution in situ hybridization (ISH) maps that allow researchers to focus on a gene of interest, and view its expression in the macaque brain across postnatal development. The dataset includes genes known to be important in brain function, "marker" genes for specific anatomical regions and cell types, disease-related genes, and genes important in brain evolution. View an example of ISH data. (pdf, 812.89 kb)
- Microarray-based profiles that reveal the expression levels of nearly all ~20,000 genes in the macaque genome. Researchers can compare genome-wide expression patterns across brain regions and ages. In addition to the five brain regions mentioned above, the data will include a set of micro-dissected brain regions for fine structure transcriptional profiling across prenatal and postnatal development for fine nuclear subdivisions of these regions. View an example of microarray data. (pdf, 162.08 kb)
- MRI scans and Nissl-stained brain slices that provide an anatomical frame of reference for the ISH and microarray data.
Contacts
Michelle Freund, Ph.D.
Chief, Neurotechnology Office of Technology Development and Coordination, NIMH
301-443-1815
Related Resources
- The Allen Institute for Brain Science built the NHP Atlas in partnership with the University of California Davis, under a contract awarded by the NIH Blueprint in September 2008. NIMH provided supplemental funds for the micro-dissection analysis.
- For technical questions about the atlas, please contact the Allen Institute.
